for Natural Philosophy of Death (2026), by Charles Dermer
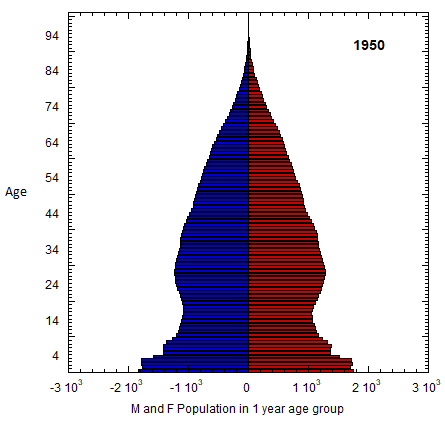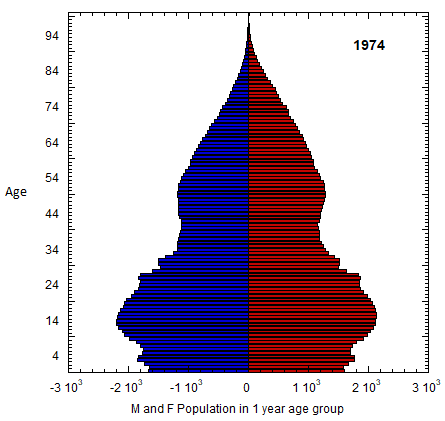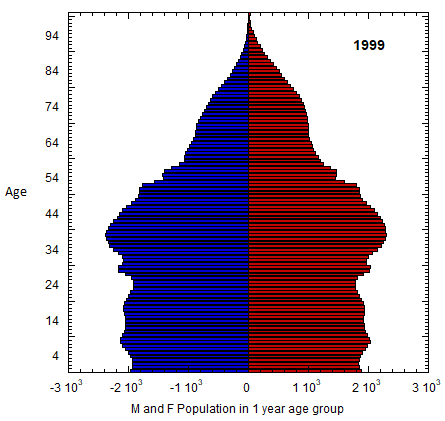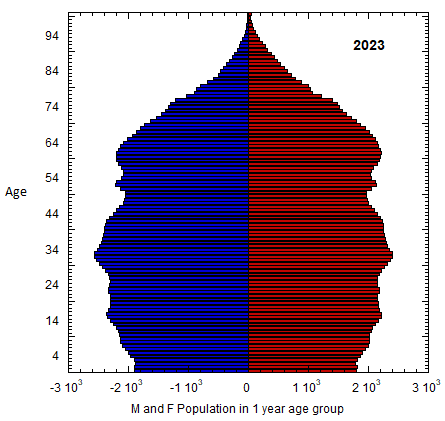
Abstract figure: US M (left) and F (right) profile evolution, 1980-2023 (populations of 1-yr age groups in thousands). The generations of Americans are defined in Table VI.C1. Changes in the number of members of co-aging age groups give the sum of the migration and mortality rates for that age group. Population rates equal mortality rates in old age. Data from USCB/WPP.
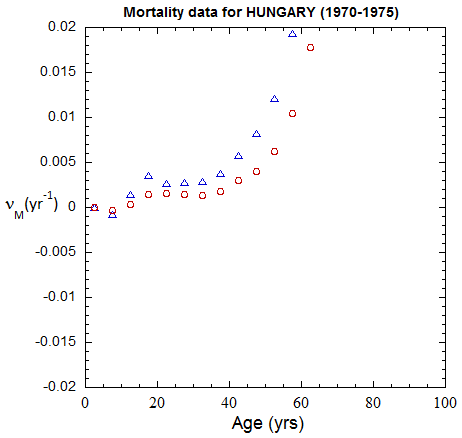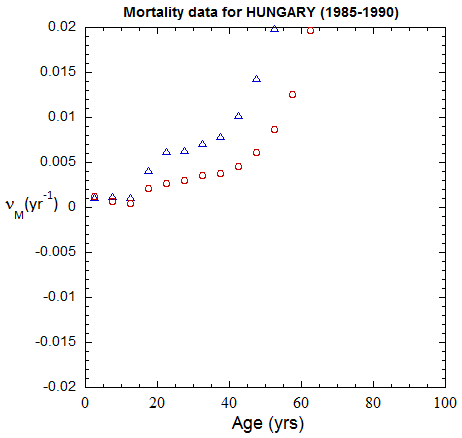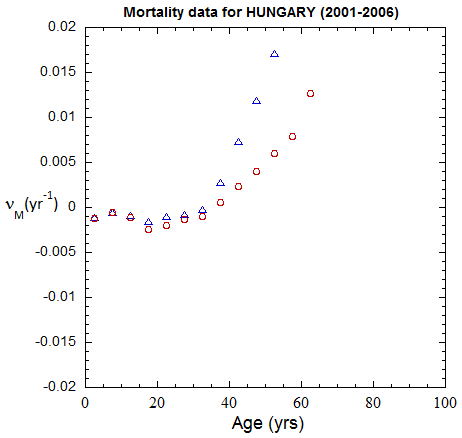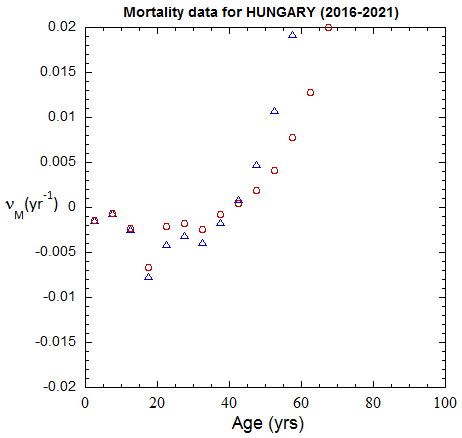
Figure II.A1a. Population rates (mislabeled “Mortality data”) rates inferred from ppn data for Hungary from 1970-2022, plotted on a linear scale. M: open diamonds. F: open circles.
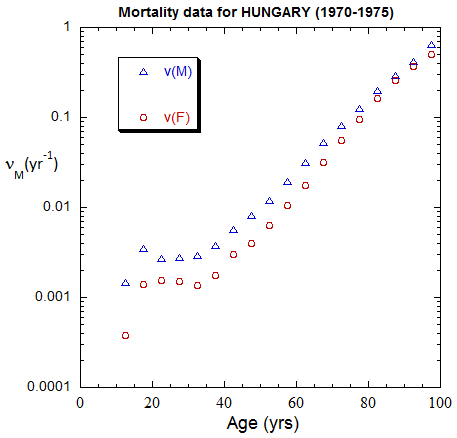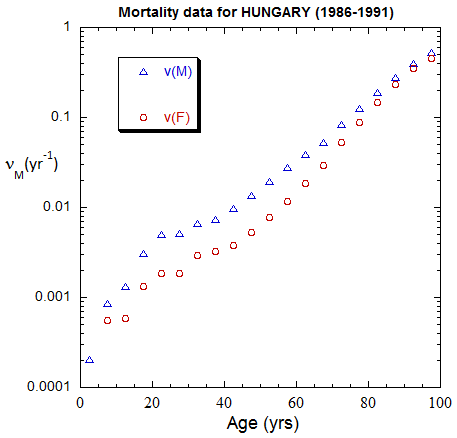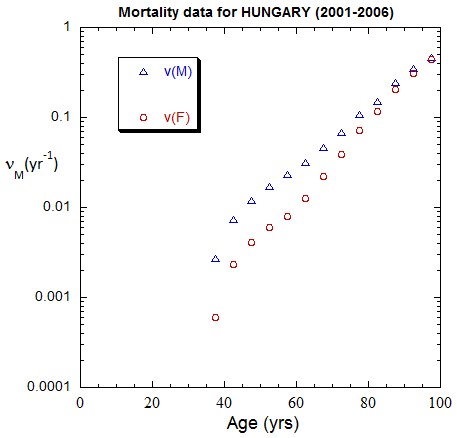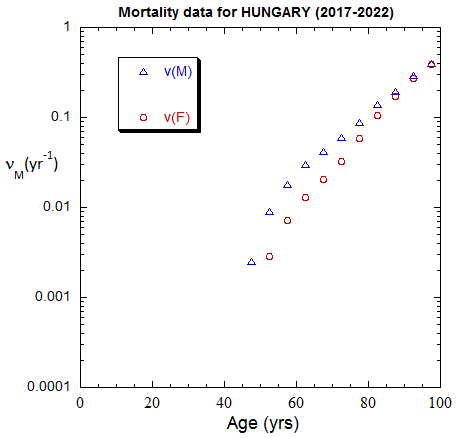
Figure II.A1b. Same as Fig. II.A1a, plotted on a logarithmic scale.
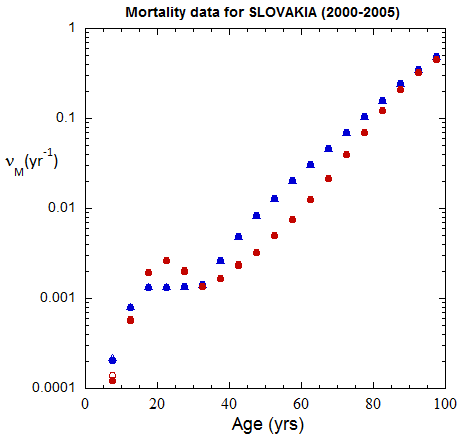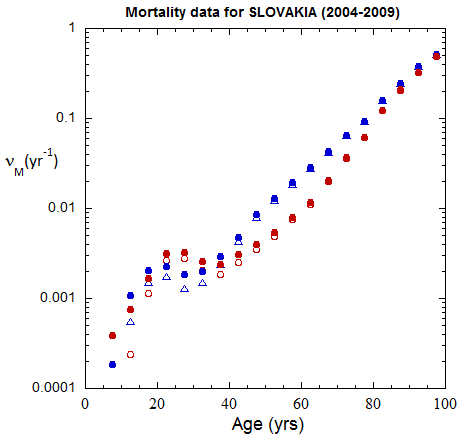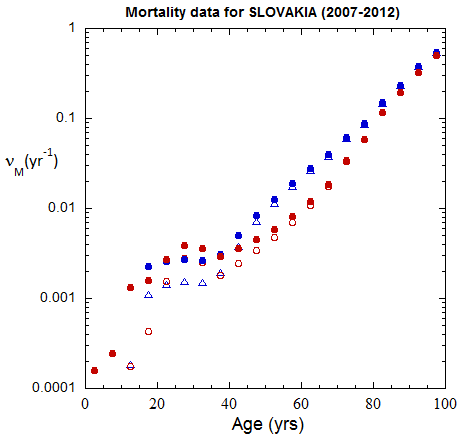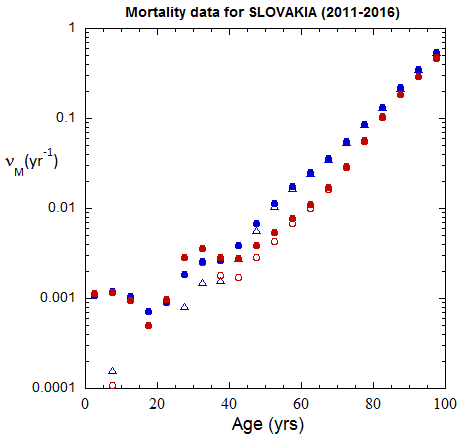
Fig II.B1. Population rates (mislabeled “Mortality data”) for Slovakia inferred from ppn data for Slovakia from 2000-2016, plotted on a logarithmic scale.
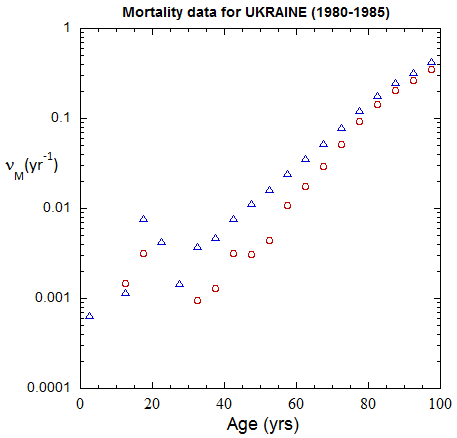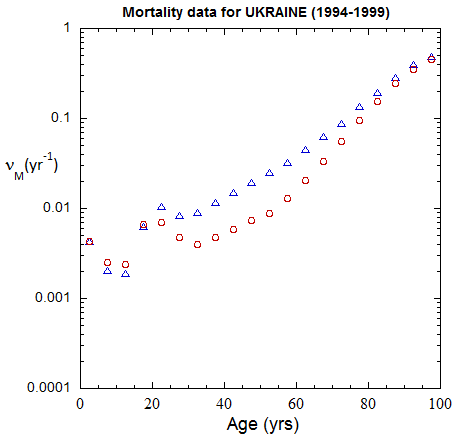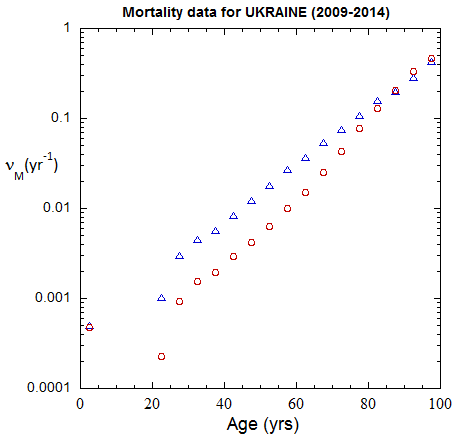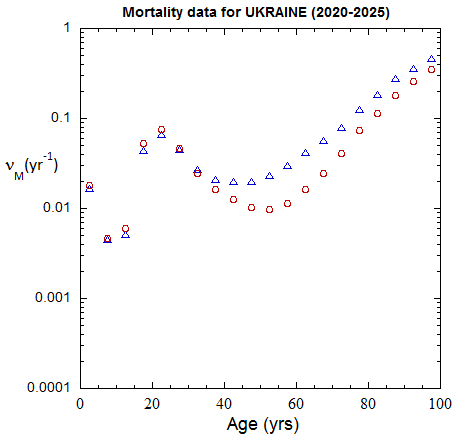
Figure III.3, III.B1. Age-stratified population rates (mislabeled “Mortality data”) derived from ppn population data for Ukrainian M (open diamonds) and F (open circles) from epoch 1980-1985 through epoch 2020-2025.
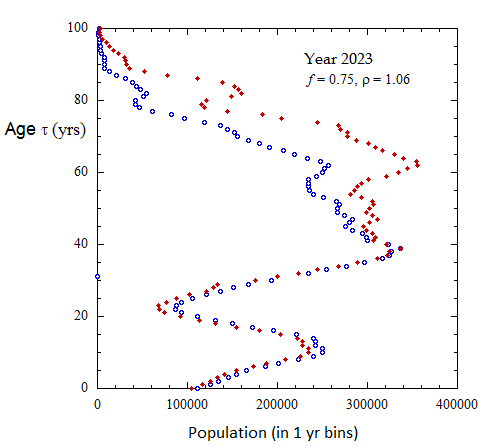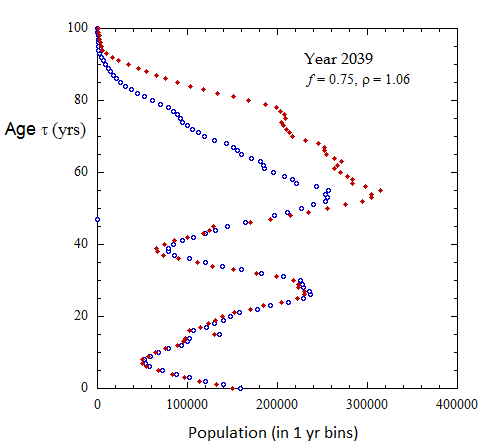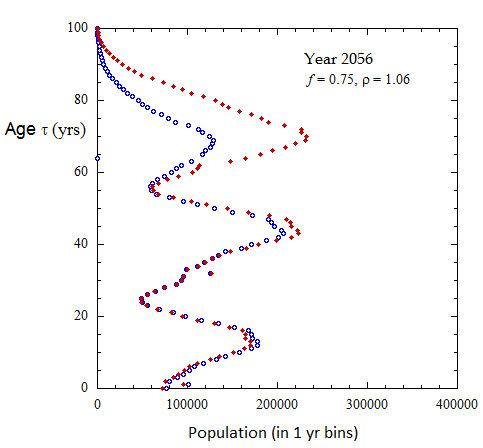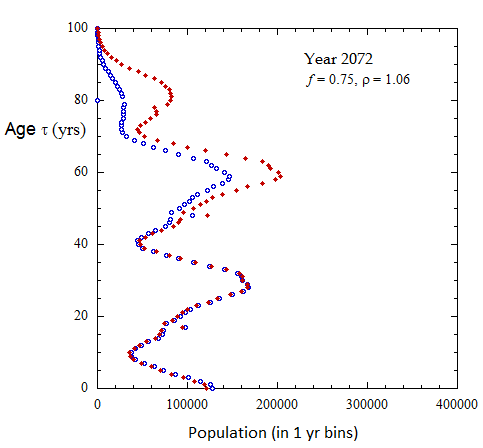
Figure III.8, III.C1. Projected Ukrainian population profiles from 2023 to 2072 for Scenario A with f = 0.75, tpk = 30 yo and birth sex ratio rho = 1.06.
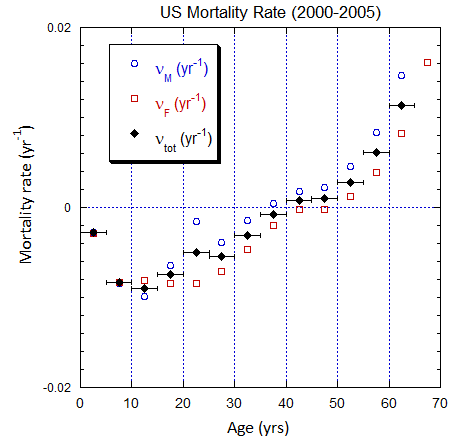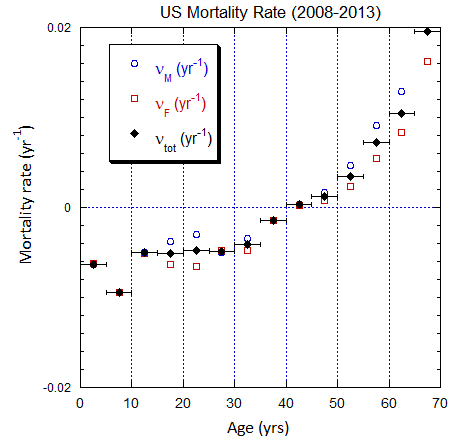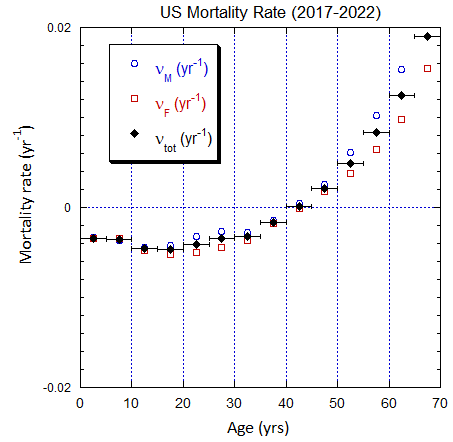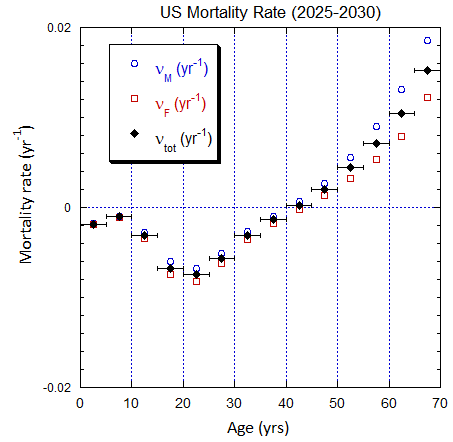
Figure V.B2. Total US population rates (mislabeled “Mortality Rate”) derived from US ppn population data from 2000-2025 (circles, M; squares, F; diamonds, average). The horizontal error bars on the average rates define the age ranges.
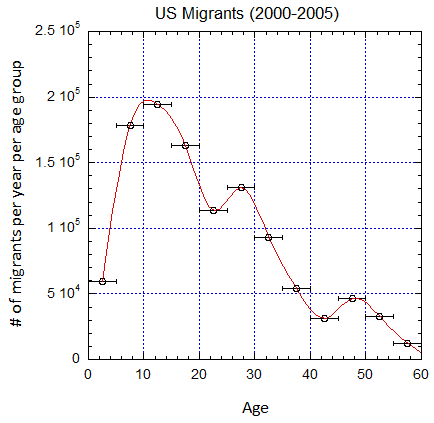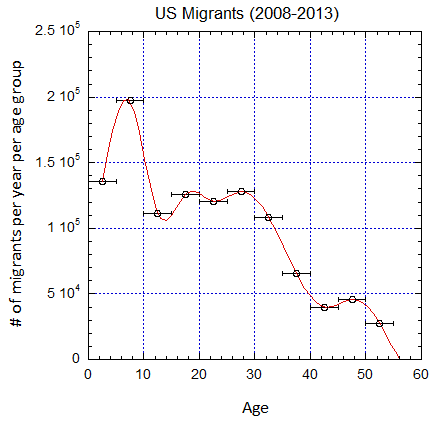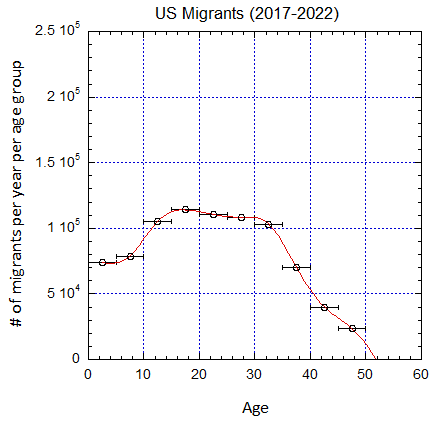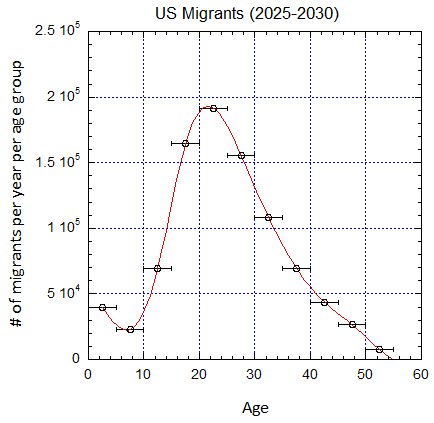
Figure V.B3. Implied number of migrants per year in the different age groups defined by the horizontal error bars from epoch 2000-2005 to 2025-2030 (UN projections for the later years) assuming a baseline 2019 mortality shown in Fig. V.B2. Cubic spline fits are shown.
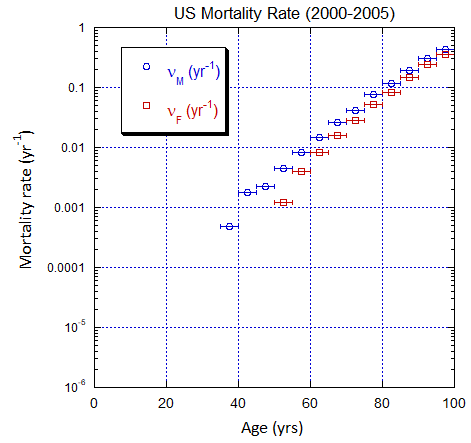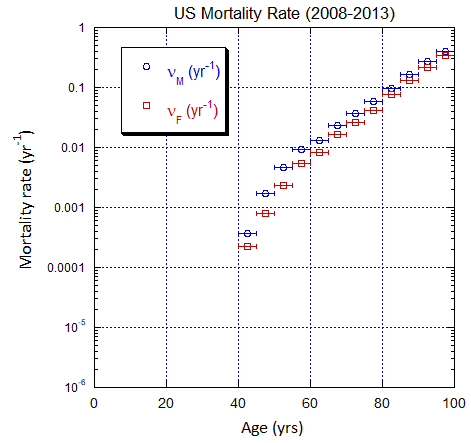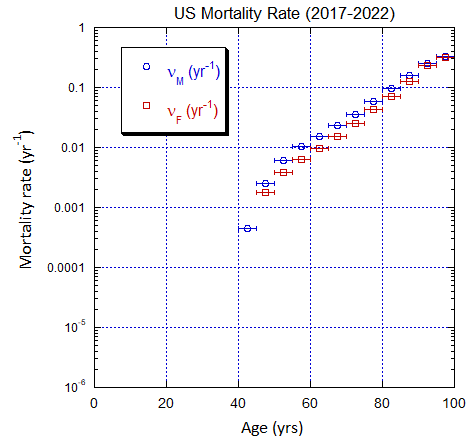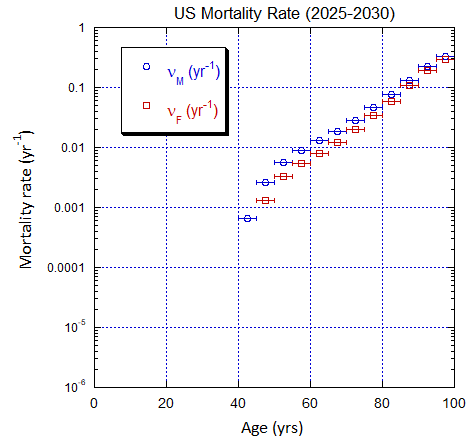
Figure V.C2. Evolving US M and F 5-yr ppn population rates (mislabeled “Mortality Rate”) averaged over the 5-yr range shown by the horizontal error bars, from epoch 2000-2005 to epoch 2025-2030. The effects of migration are large for ages ≲50 yo. The rates in old age approach mortality rates. See Fig. VI.22 for Loch Ness animation.
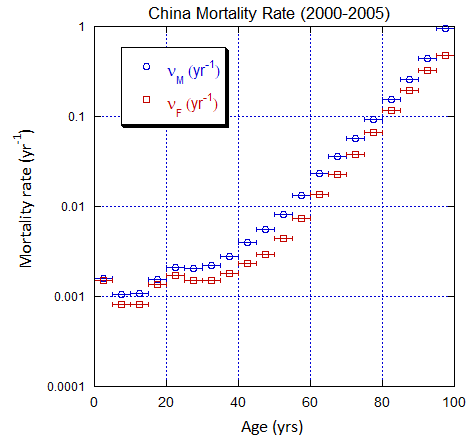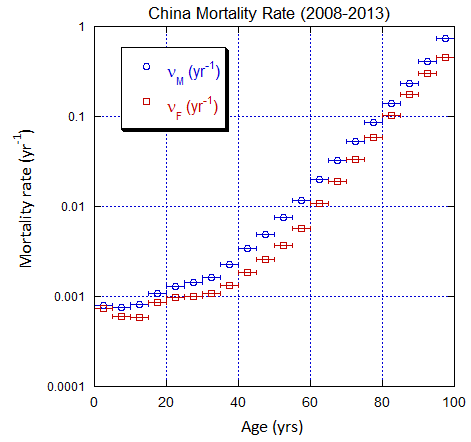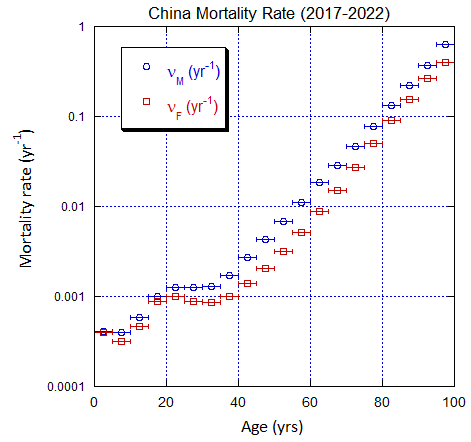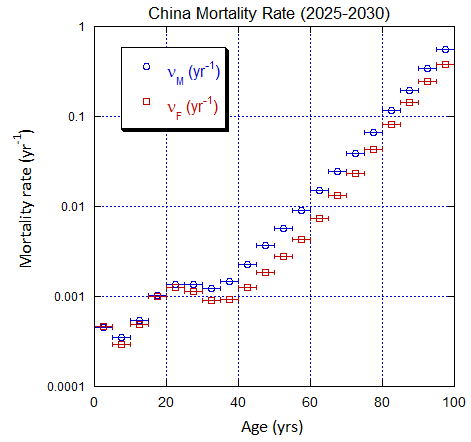
Figure V.C3. Evolving China M and F age-stratified population rates derived from ppn data for China. Note the regular behavior. Open circles: M, open squares: F.
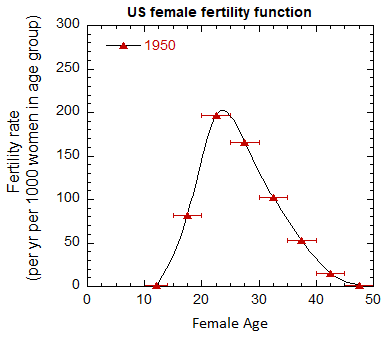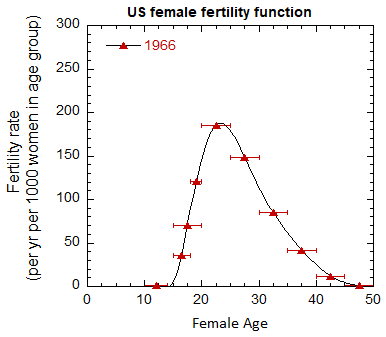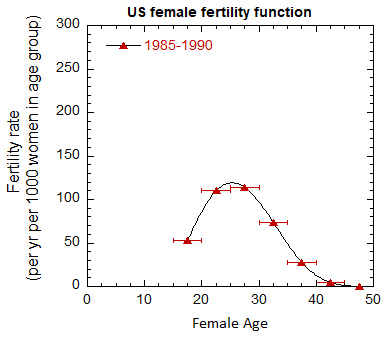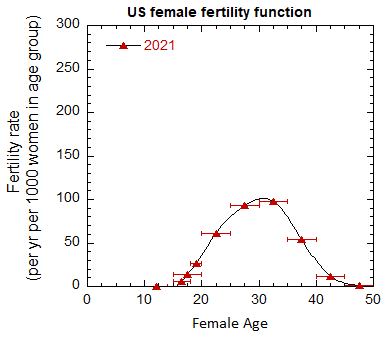
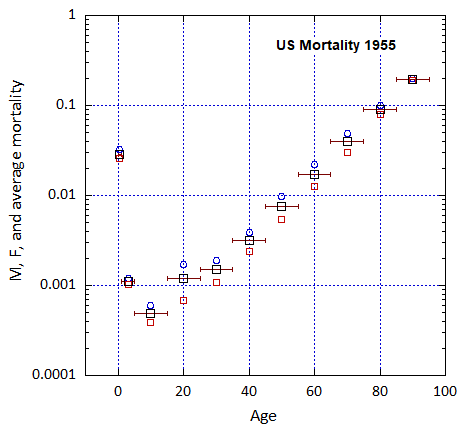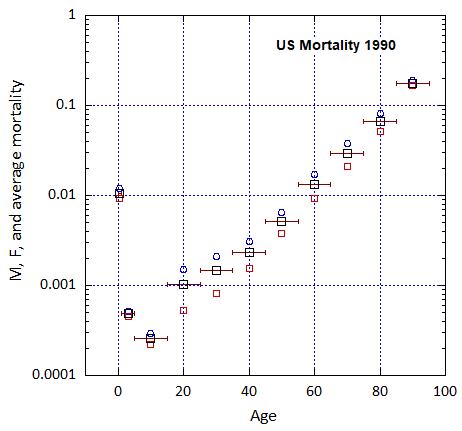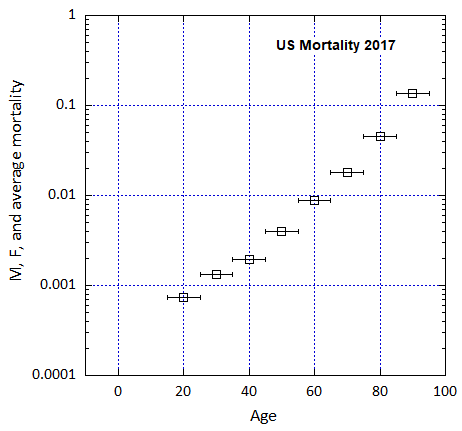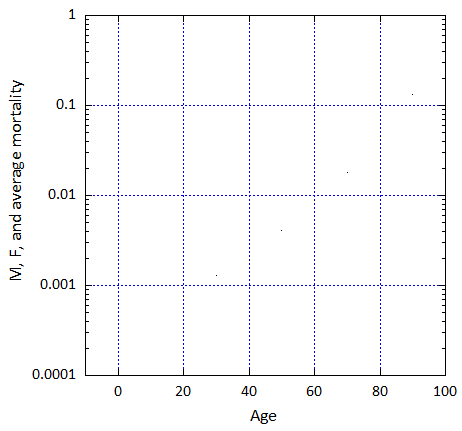
Figure VI.5. Total US M, F and average M+F mortality rates from 1955-2022.
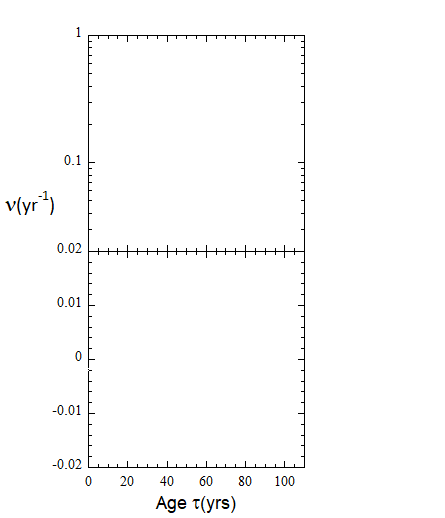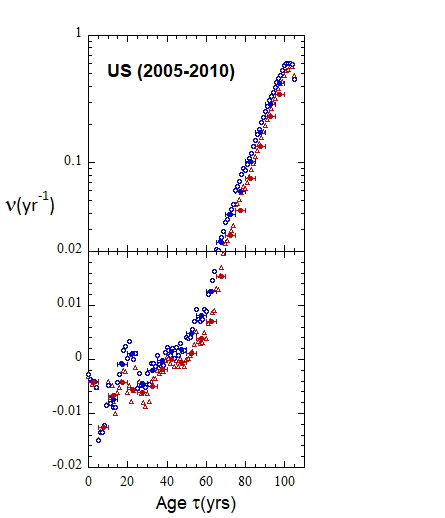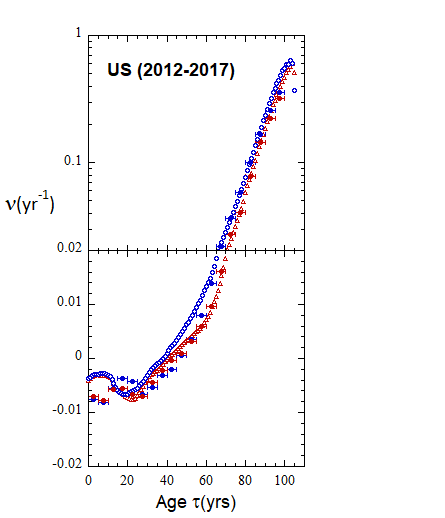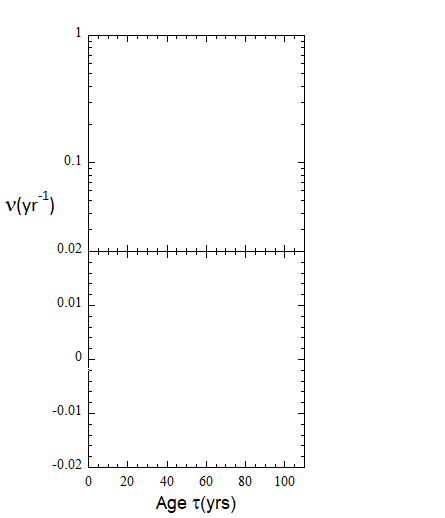
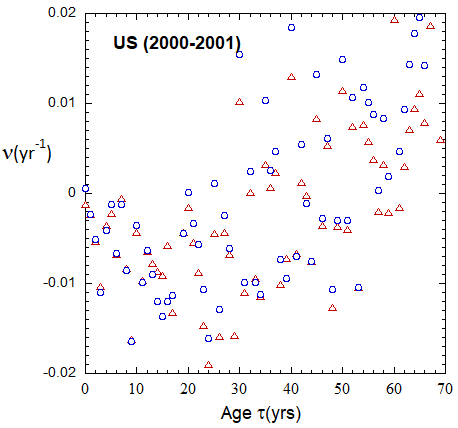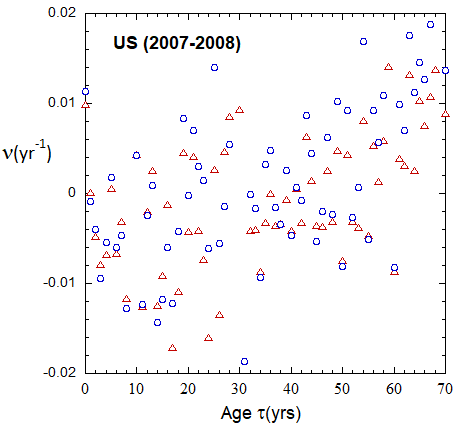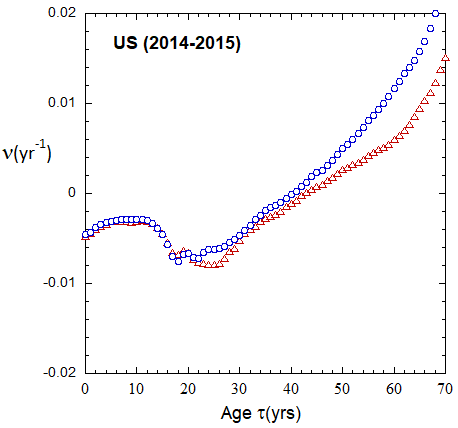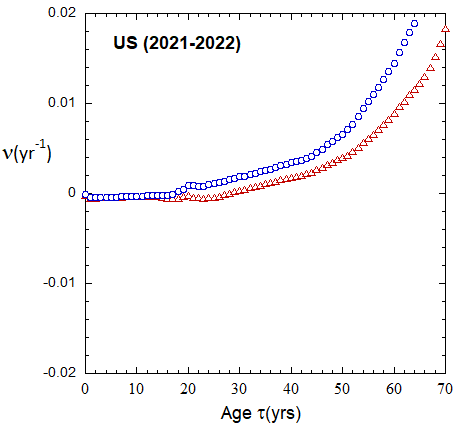
Figure VI.19a. Animations of 1-yr US M and F rates from 2000 to 2021 derived from 1-yr age cohort data given in the HMD. Rates plotted on a linear scale.
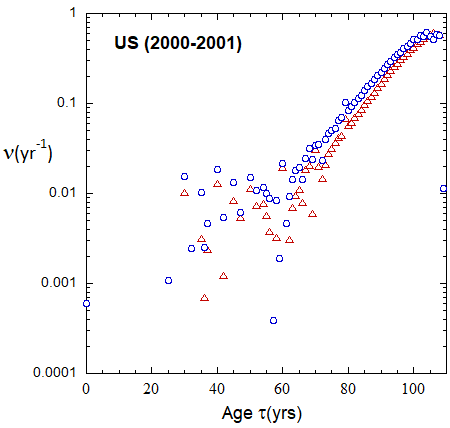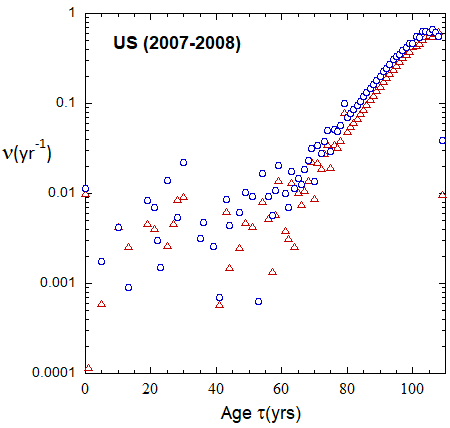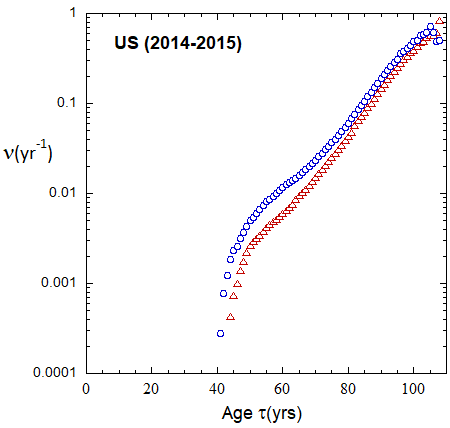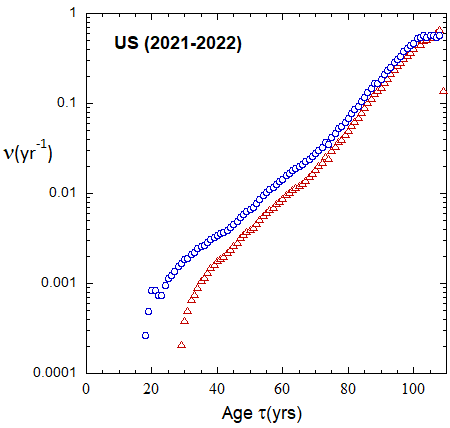
Figure 19b: As in FIg. 19a, but with rates plotted on a logarithmic scale.
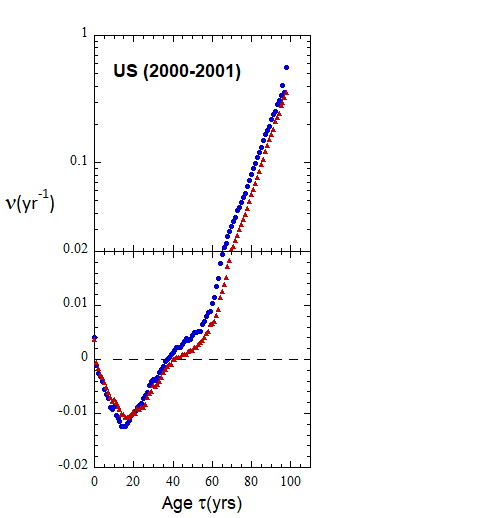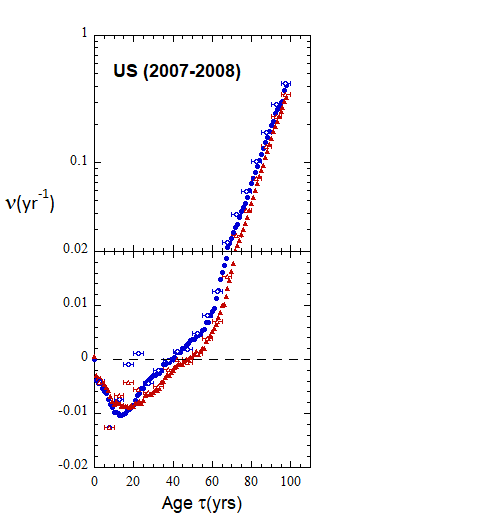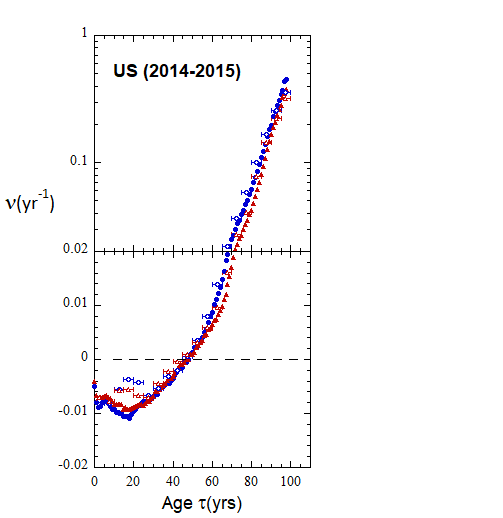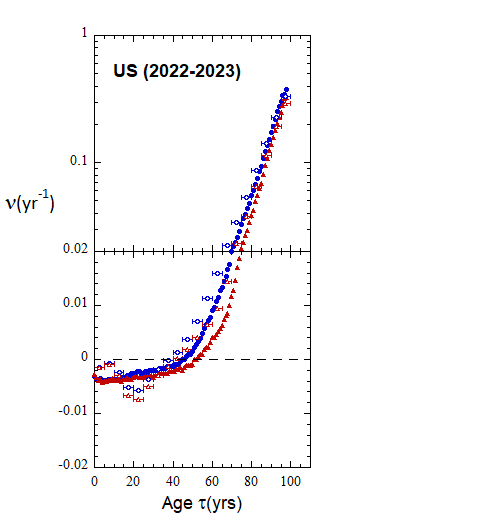
Figure VI.22. Animation of Loch Ness plots for 1-yr rates derived for the US from epoch 2000-2010 to epoch 2022-2023 from WPP/USCB data. Rates from 5-yr age-cohort (ppn) data are shown for comparison.
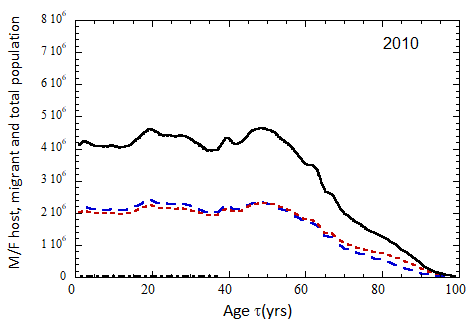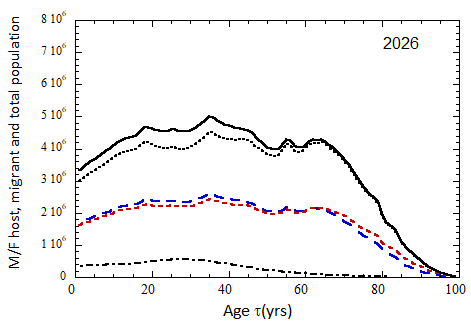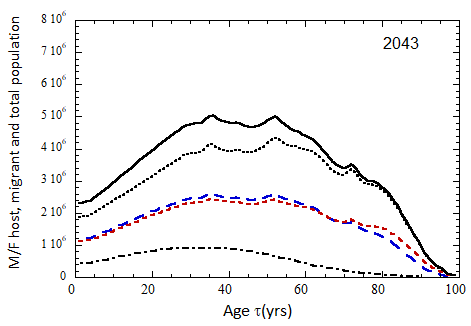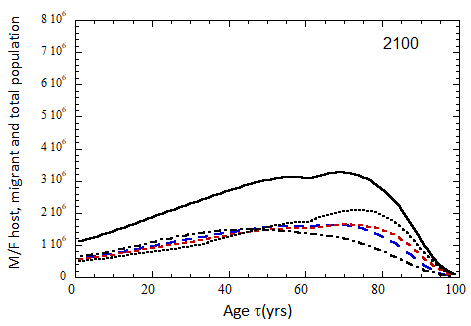
Figure VII.15a. Host (dotted), migrant (dot-dashed) and host + migrant (solid) total (M + F) profiles for the 2010 M (dashed) and F (short-dashed) profiles evolved to the year 2100. TFR model (a) is used for fertility and Fit 3 for mortality. Only migrants entering the US since 2010 are followed, and the migrant arrival rate is assumed to be equal to 1.18×106 migrants/yr after 2023
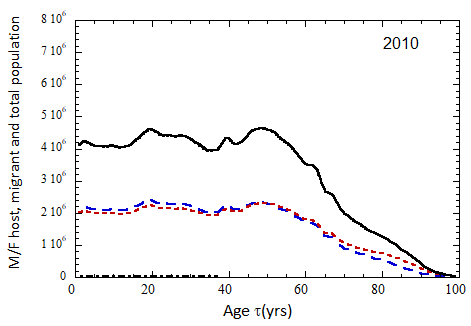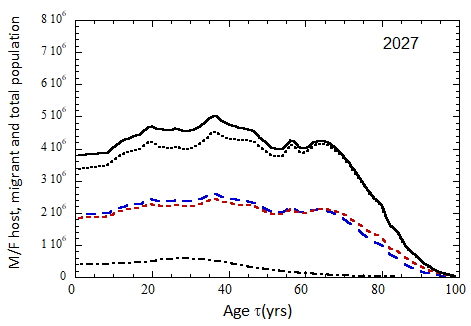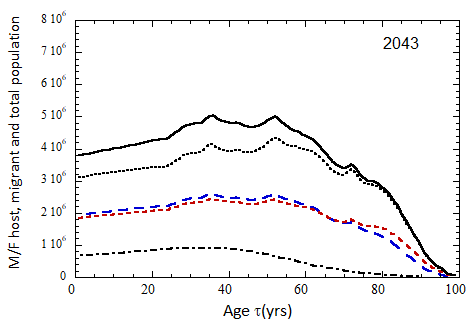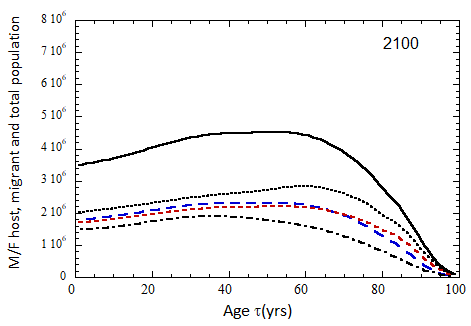
Figure VII.15b. Same as Fig VII.15a, except TFR model (b) is used for fertility.
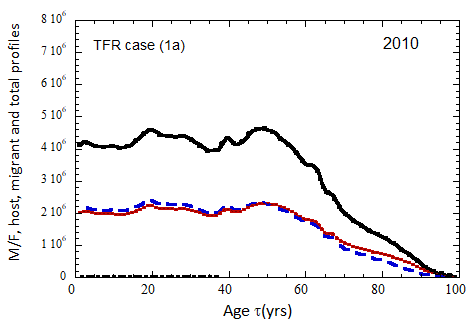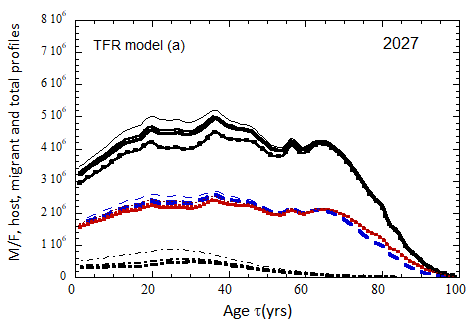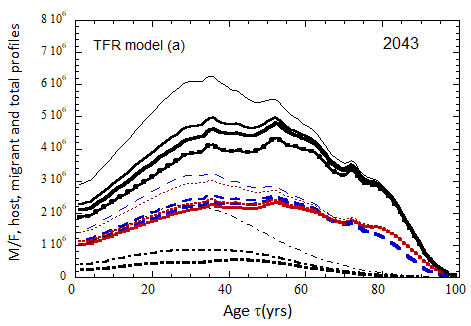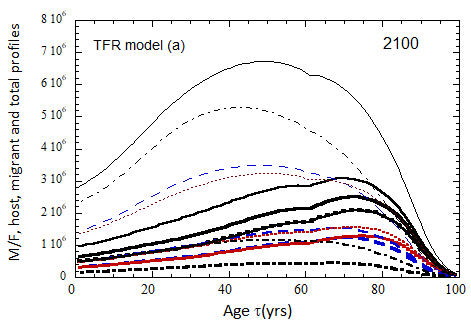
Figure VII.18. Animation of the evolution of the US M+F total population (solid curves), host population (braided curve), and migrant population (dot-dashed curves) from 2010 to 2100 starting from the USCB 2010 data. A migrant population is assumed to enter the US at the historical rate (Fig. VI.1b) prior to 2023, and at a rate equal to 0.1% (heavy curves), 0.3% (medium curves), and 1.0% (light curves) of the total population in 2024 and thereafter. Total (migrant + host) profiles for M (dashed) and F (dotted) are also shown.
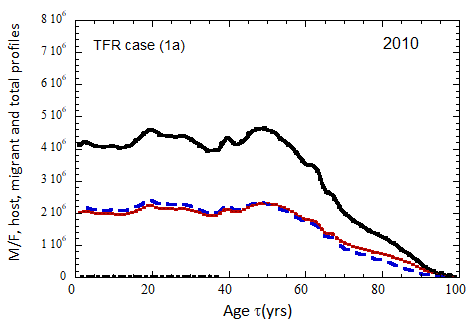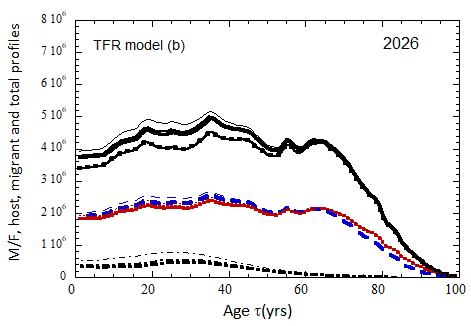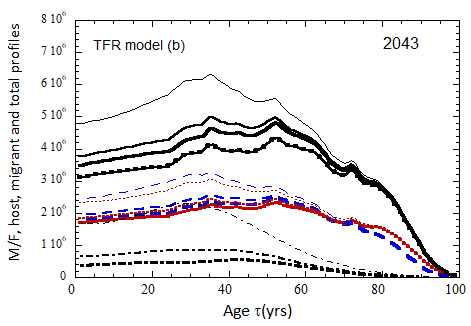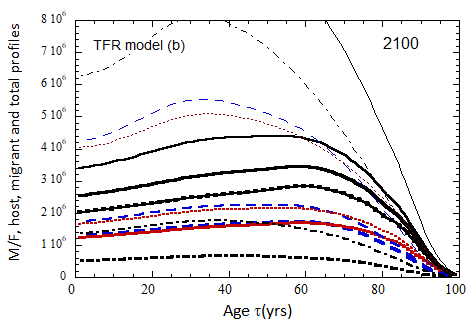
Fig. VII.18a. (above left) TFR model (a) for both host and migrant populations.
Fig VII.18b. (left) As in Fig. VII.18a but with TFR model (b) for both host and migrant populations.
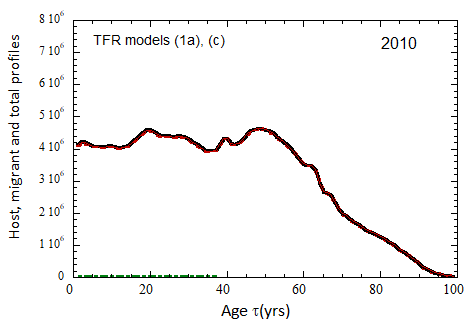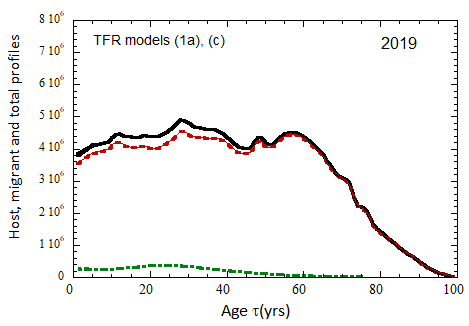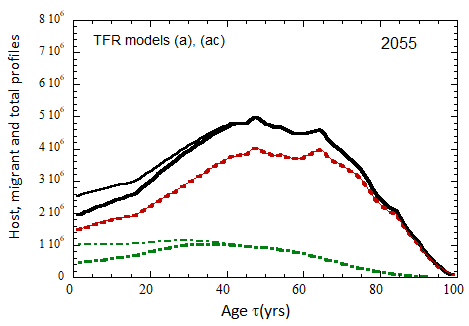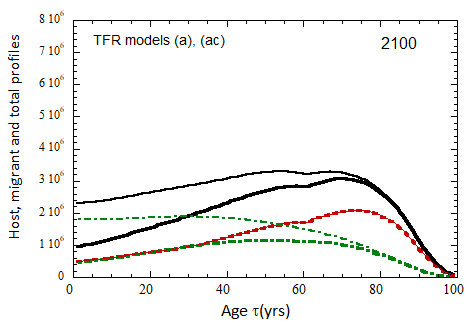
Figure VII.20a. US profile evolution starting from 2010 USCB population data and continuing to 2100. The profiles are evolved to 2020 with migration rates, mortalities, and fertilities as described in the text. After 2020, thick curves show the results when both host and migrant populations follow TFR model (a). Thin curves show results when host population follows TFR model (a) and the migrant population is assumed to have TFR = 2.0. After 2023, the migration-rate factor fm= 0.3, that is, the migrant arrival rate into the US is assumed to be 0.3% of the total host + migrant population in that year. Braided curve: host population evolution. Dot-dashed curves: migrant age distribution since 2010. Solid curves: age distributions of host + migrant population.
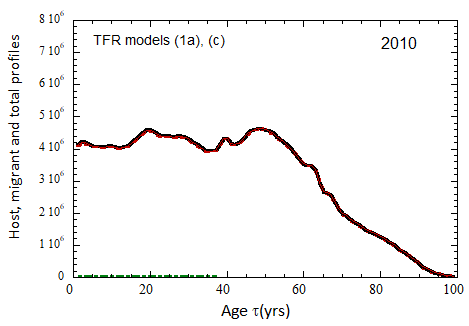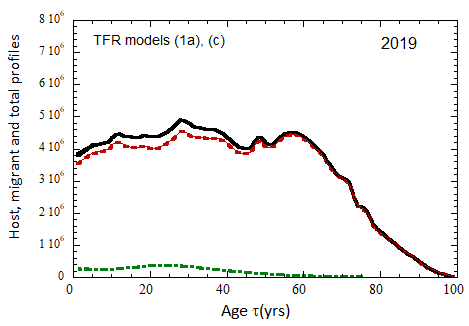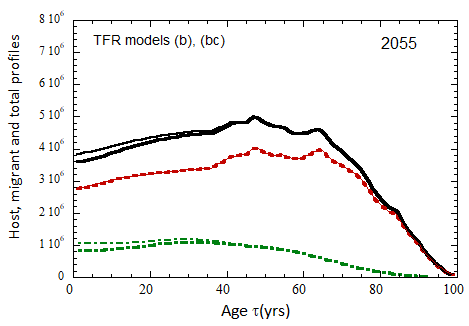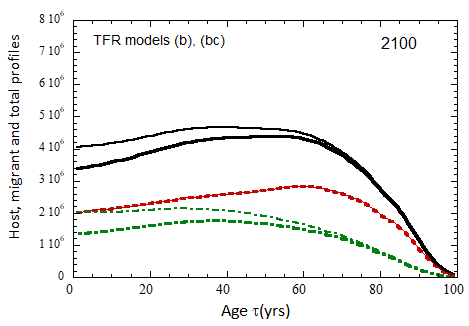
Figure VII.20b. Same as Fig. VII.20a, except that after 2020, thick curves show the results when both host and migrant populations follow TFR model (b). Thin curves show results when host population follows TFR model (b) and the migrant population is assumed to have TFR = 2.0.
Statistical Mechanics Simulations
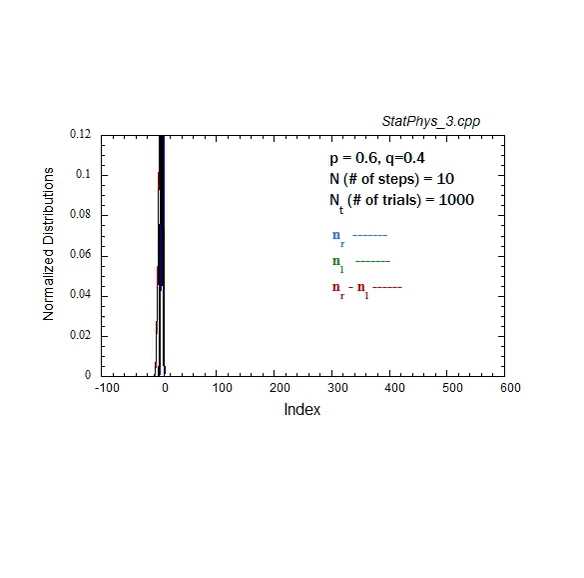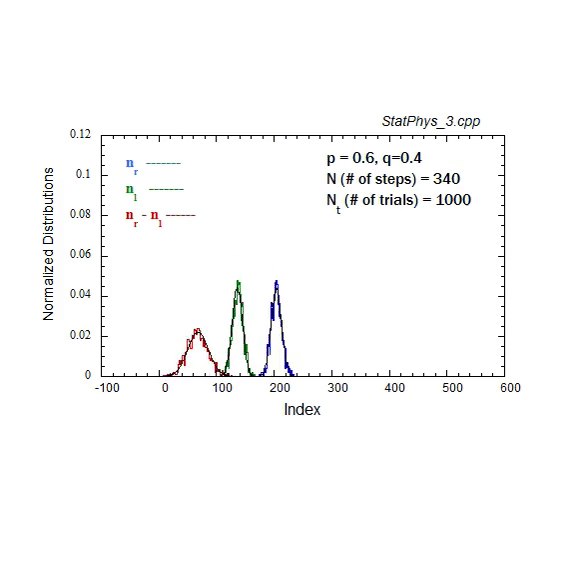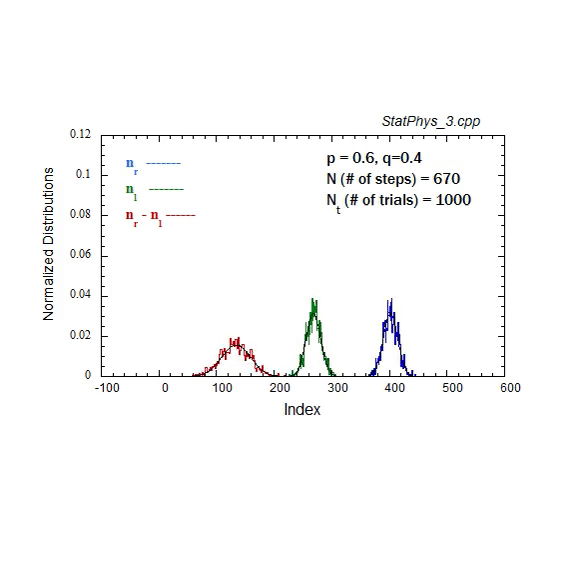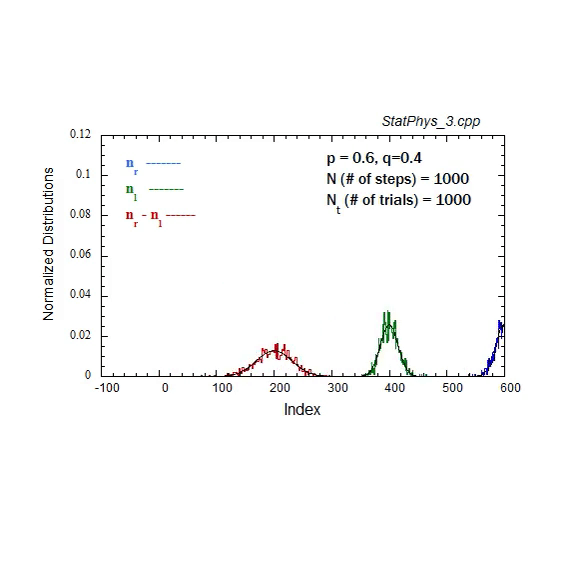
Figure A1. Formation and evolution of Gaussians from weighted binomial distribution for steps to right and left, and net displacement.
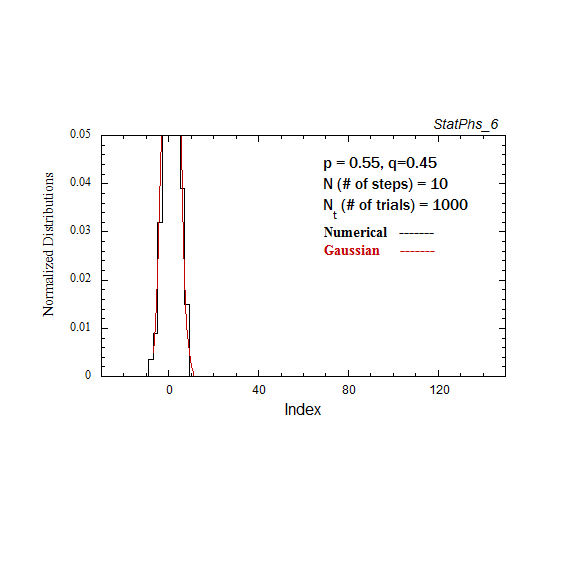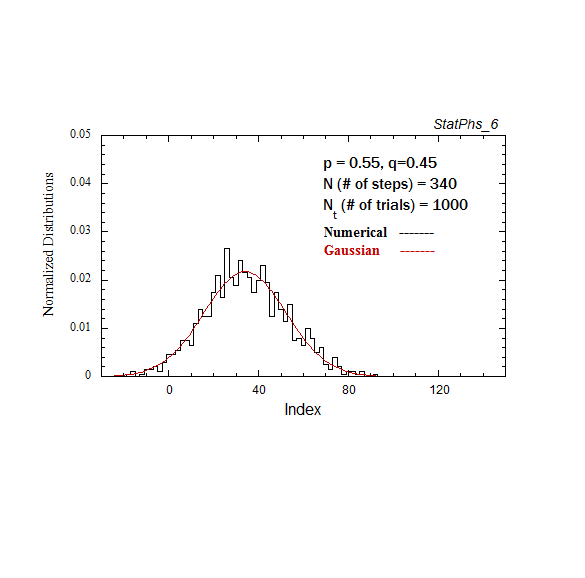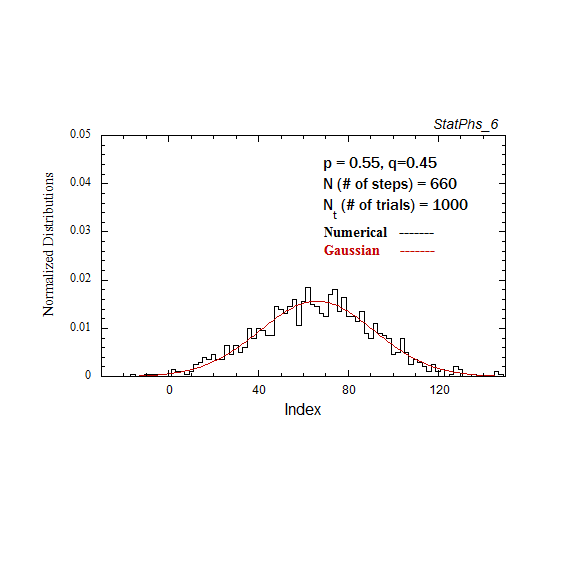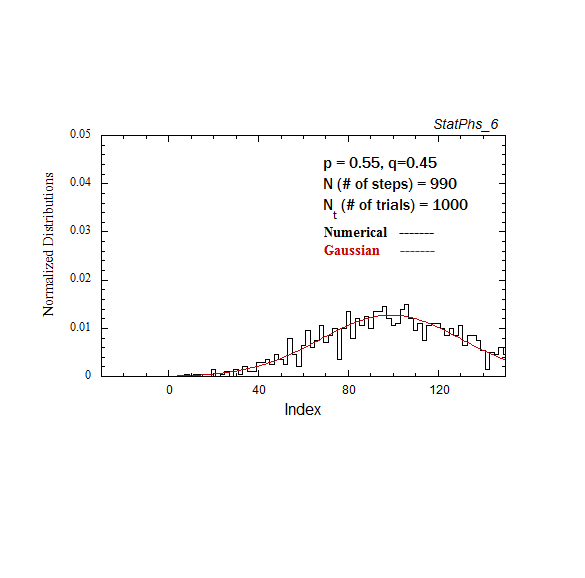
Figure A2. Particle distribution evolving in time, with p = 0.55. The smooth solid curve is the analytic expression.
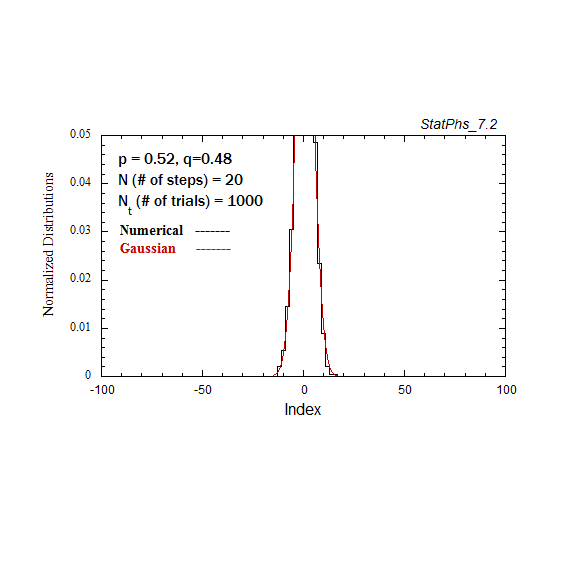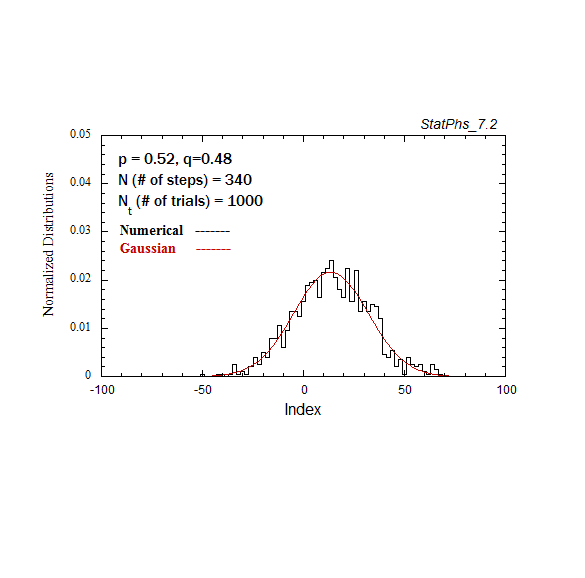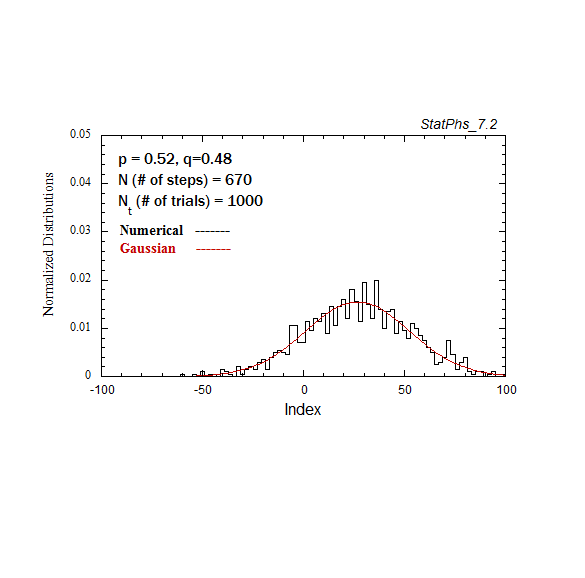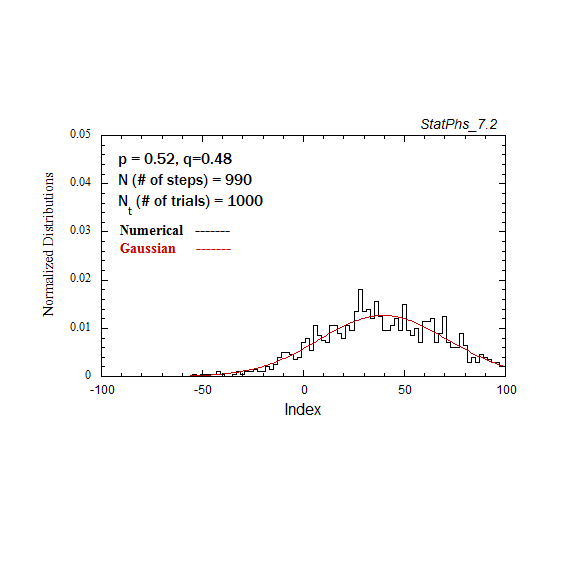
Figure A2a. Particle distribution evolving in time as in Fig. A2, but with p = 0.52.
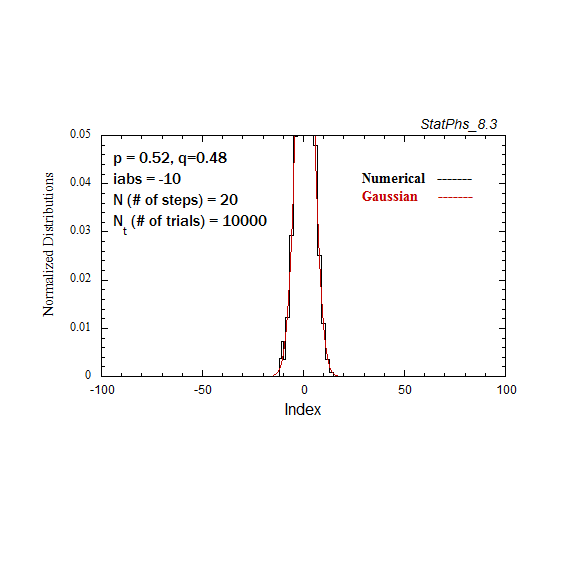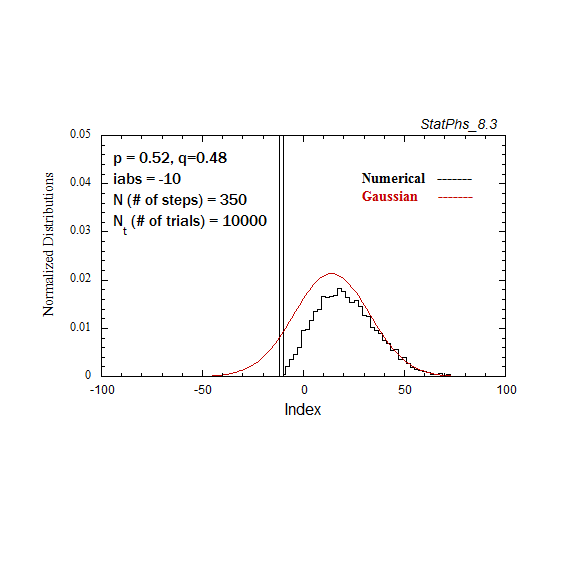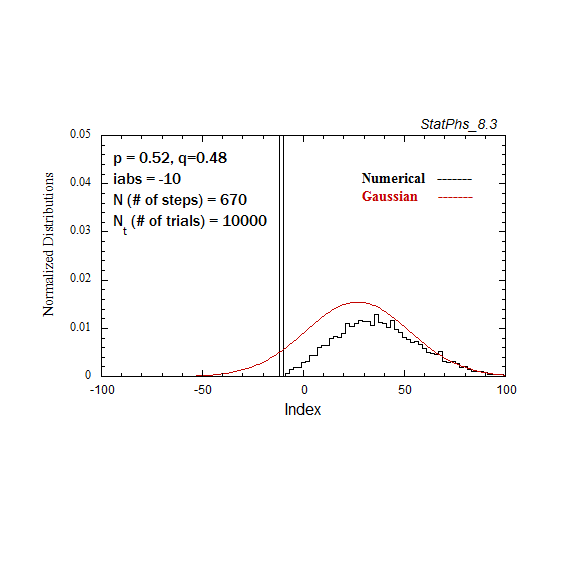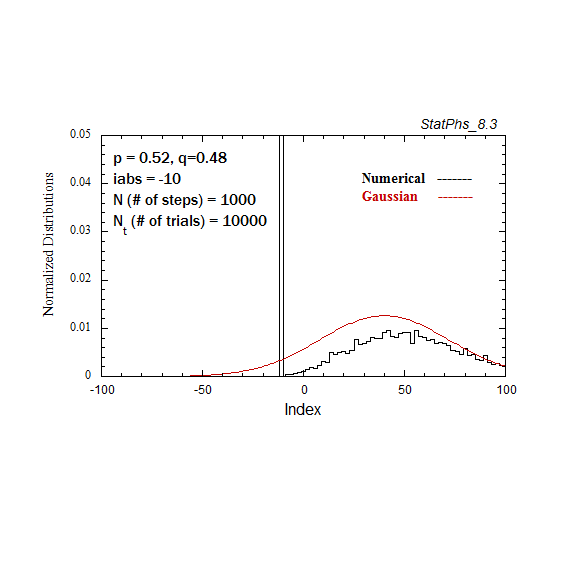
Figure A3. Evolving particle distribution in presence of an absorbing boundary, with p=0.52.
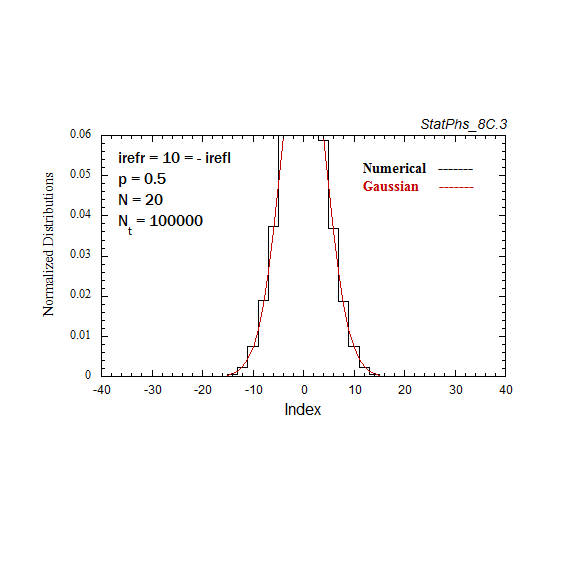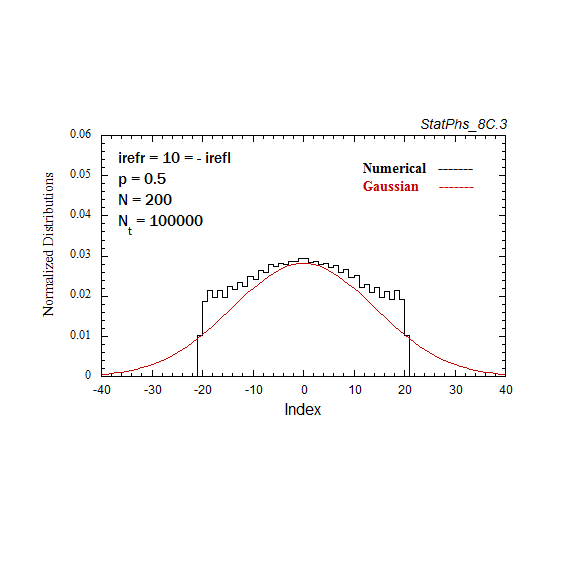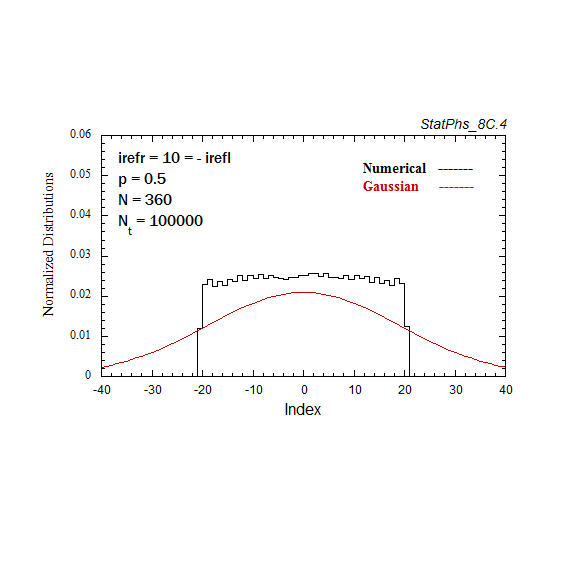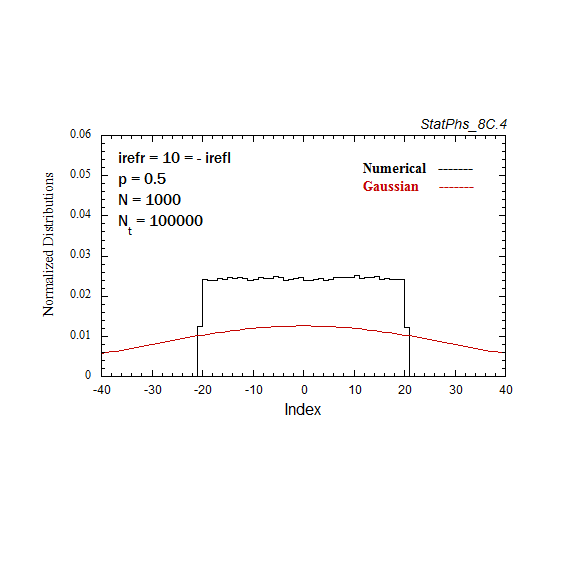
Figure A4. Evolving particle distribution in presence of two reflecting boundaries at -20 and +20.
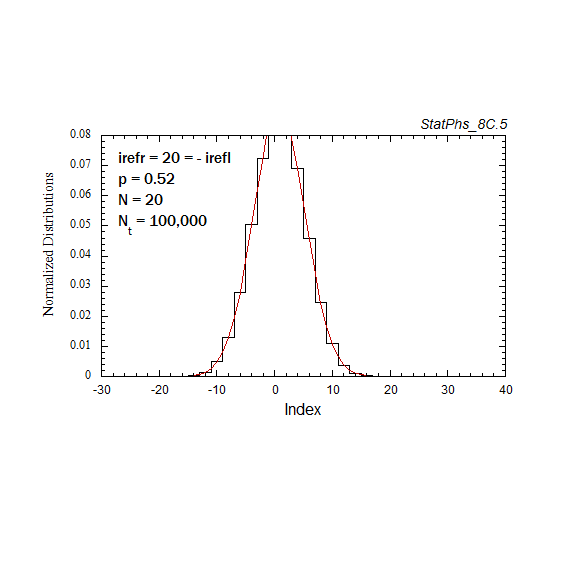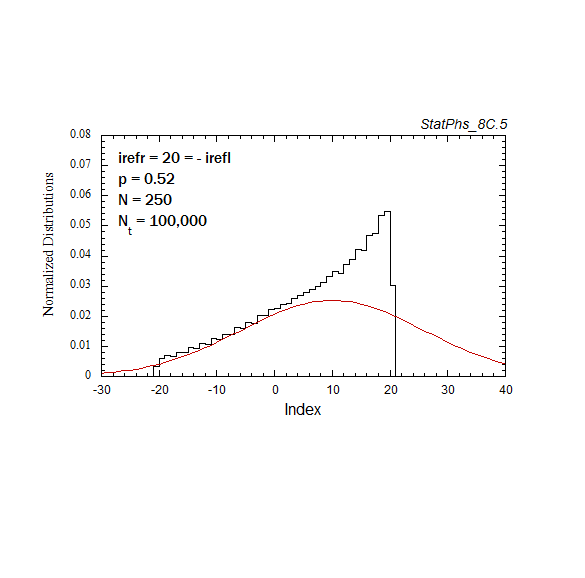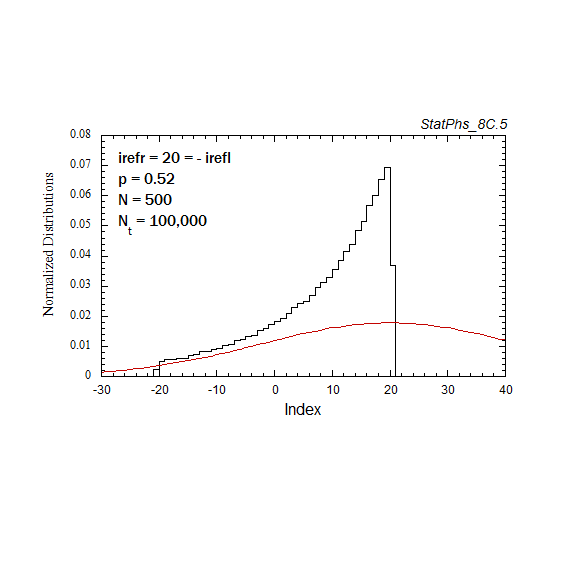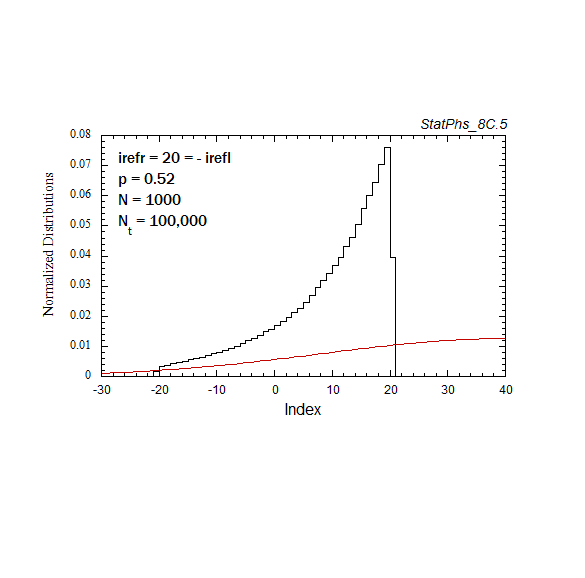
Figure A6. Evolution of particle distribution with reflecting boundaries and different step probabilities to the left and right.
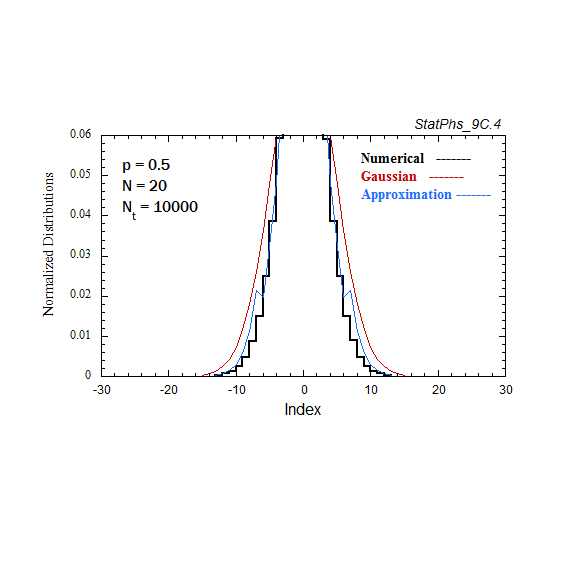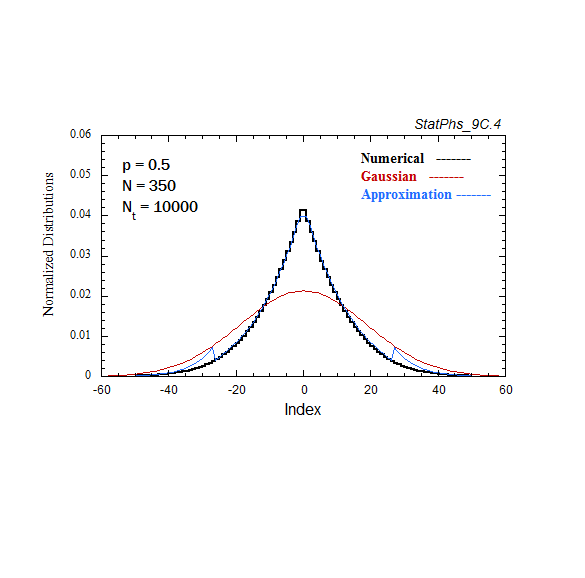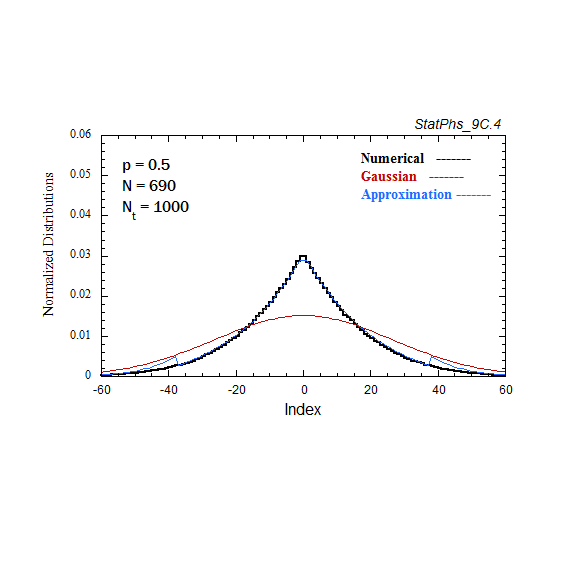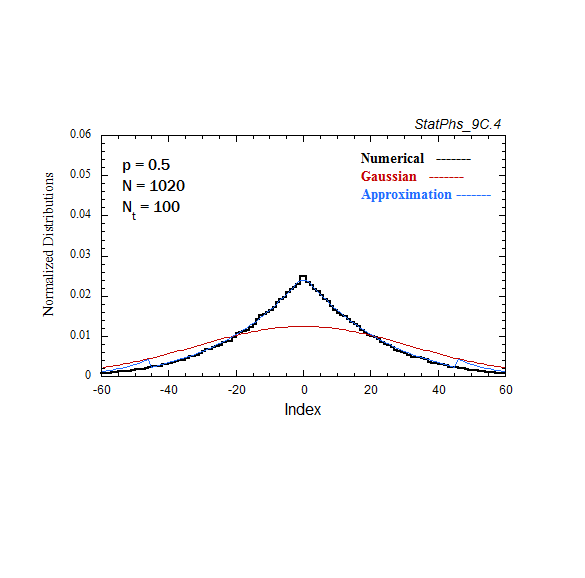
Figure A7. Comparison of Gaussian and analytic approximations of evolving particle distributions with continuous injection. The total number of injected particles is renormalized to the same number of particles as in the evolving Gaussian. Two asymptotes of the analytic approximation are shown.

Figure A8a. Evolution of a particle distribution with continuous injection, absorbing and reflecting boundaries, and step probabilities as noted. Gaussian and continuous injection approximations in the absence of boundaries are shown for comparison. The distributions are normalized to unity.
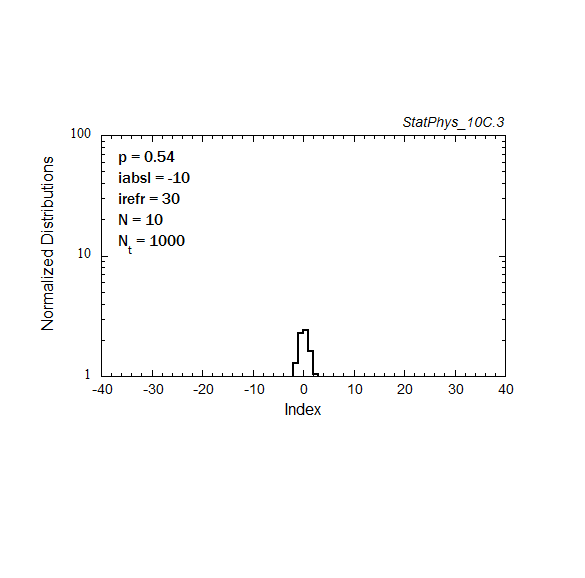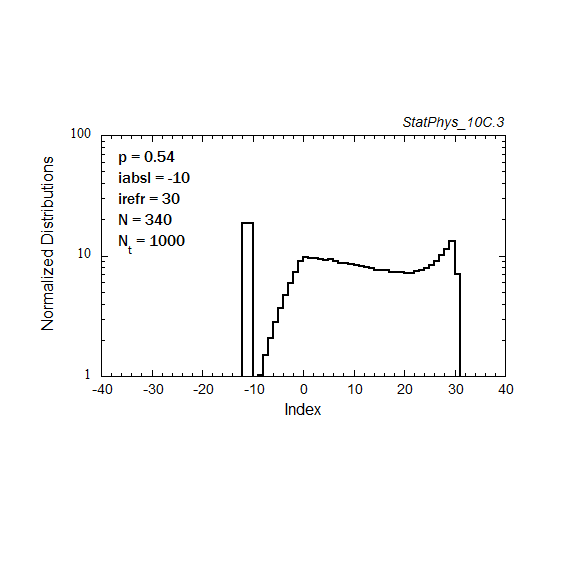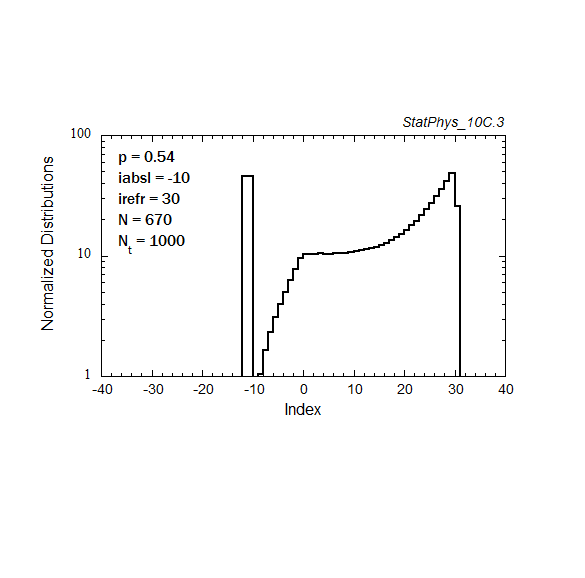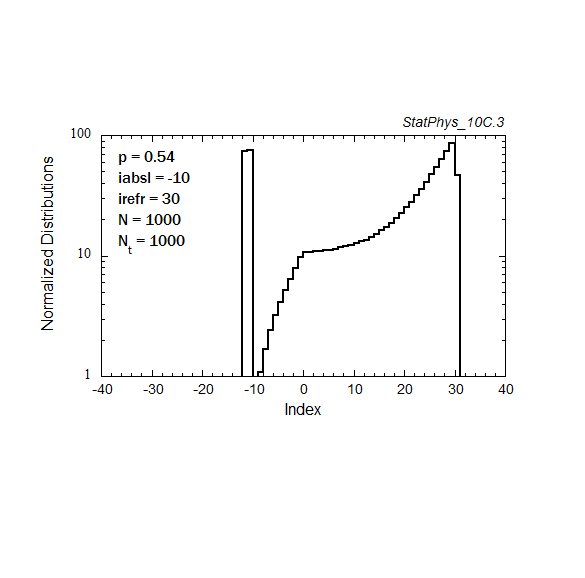
Figure A8b. Same as Fig. A8a, but plotted on a logarithmic scale for the distribution, with absolute normalization to total number of injected particles.
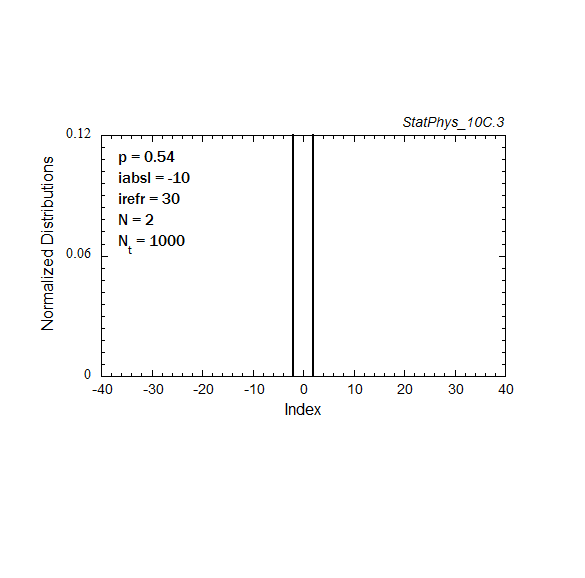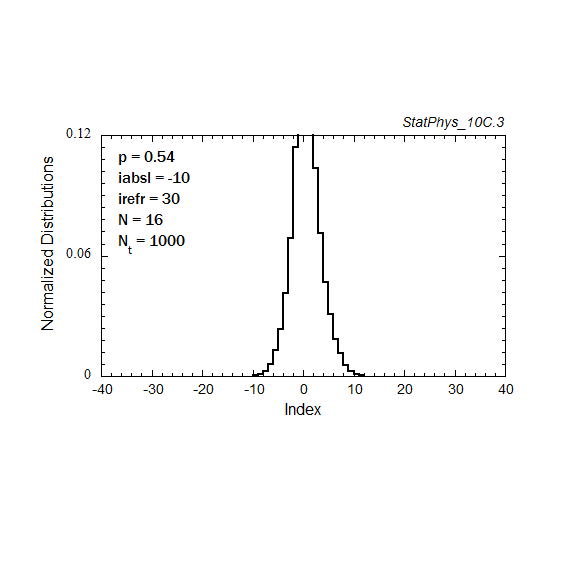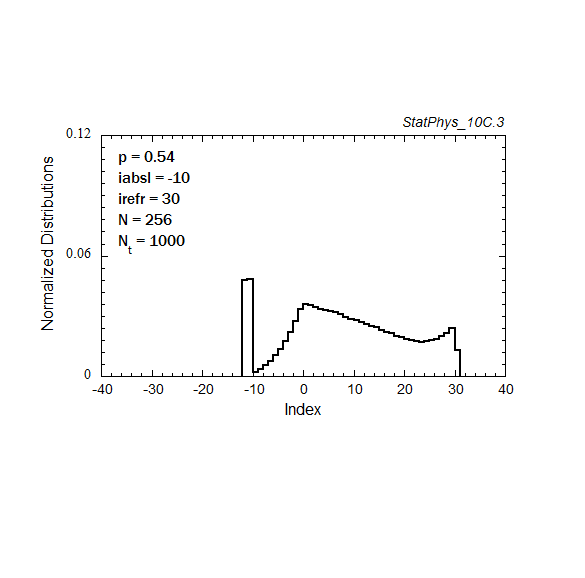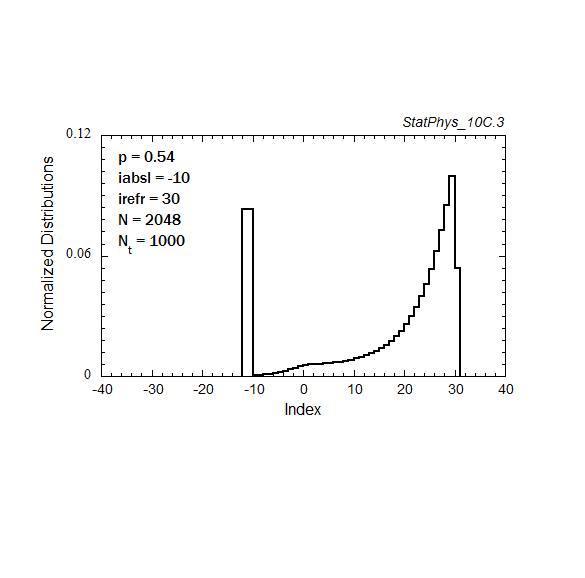
Figure A8c. Same as Fig. A8a, but with logarithmic time spacings.